Our primary goal is to better understand how biological molecules behave in space and time.


Atsushi Miyawaki, M.D., Ph.D.
Team Director, Cell Function Dynamics
miyawaki-lab.ast@ml.riken.jp
Research Overview
The behavior of biochemical molecules moving around in cells makes me think of a school of whales wandering in the ocean, captured by the Argos system on the artificial satellite. When bringing a whale back into the sea --- with a transmitter on its dorsal fin, every staff member hopes that it will return safely to a school of its species. A transmitter is now minute in size, but it was not this way before. There used to be some concern that a whale fitted with a transmitter could be given the cold shoulder and thus ostracized by other whales for “wearing something annoying.” How is whale’s wandering related to the tide or a shoal of small fish? What kind of interaction is there among different species of whales? We human beings have attempted to fully understand this fellow creature in the sea both during and since the age of whale fishing. In a live cell imaging experiment, a luminescent probe replaces a transmitter. We label a luminescent probe on a specific region of a biological molecule and bring it back into a cell. We can then visualize how the biological molecule behaves in response to external stimulation. Since luminescence is a physical phenomenon, we can extract various kinds of information by making full use of its characteristics. Cruising inside cells in a supermicro corps, gliding down in a microtubule like a roller coaster, pushing our ways through a jungle of chromatin while hoisting a flag of nuclear localization signal --- we are reminded to retain a playful and adventurous perspective at all times. What matters is mobilizing all capabilities of science and giving full play to our imagination. We believe that such serendipitous findings can arise out of such a sportive mind, a frame of mind that prevails when enjoying whale-watching.
Main Research Fields
Biological Sciences
Related Research Fields
Chemistry / Biophysics
Keywords
- Bioimaging
- Fluorescence protein
- Chromophore
Selected Publications
- Ando R., Shimozono S., Ago H., Takagi M., Sugiyama M., Kurokawa H., Hirano M., Niino Y., Ueno G., Ishidate F., Fujiwara T., Okada Y., Yamamoto M., Miyawaki A.:
“StayGold variants for molecular fusion and membrane-targeting applications.”
Nature Methods 21(4):648-656. (2024) - Hirano M., Ando R., Shimozono S., Sugiyama M., Takeda N., Kurokawa H., Deguchi R., Endo K., Haga K., Takai-Todaka R., Inaura S., Matsumura Y., Hama H., Okada Y., Fujiwara T., Morimoto T., Katayama K., Miyawaki A.:
“A highly photostable and bright green fluorescent protein.”
Nature Biotechnology 40:1132-1142. (2022) - Michikawa T., Yoshida T., Kuroki S., Ishikawa T., Kakei S., Kimizuka R., Saito A., Yokota H., Shimizu A., Itohara S., Miyawaki A.:
“Distributed sensory coding by cerebellar complex spikes in units of cortical segments.”
Cell Reports 37(6):109966. (2021) - Katayama H., Hama H., Nagasawa K., Kurokawa H., Sugiyama M., Ando R., Funata M., Yoshida N., Homma M., Nishimura T., Takahashi M., Ishida Y., Hioki H., Tsujihata Y., Miyawaki A.:
“Visualizing and modulating mitophagy for therapeutic studies of neurodegeneration.”
Cell 181: 1176-1187 (2020) - Iwano S., Sugiyama M., Hama H., Watakabe A., Hasegawa N., Kuchimaru T., Tanaka K. Z., Takahashi M., Ishida Y., Hata J., Shimozono S., Namiki K., Fukano T., Kiyama M., Okano, S., Kizaka-Kondoh H., McHugh, T.J., Yamamori T., Hioki H., Maki S., Miyawaki A.:
“Single-cell bioluminescence imaging of deep tissue in freely moving animals.”
Science 359: 935-939. (2018) - Sakaue-Sawano A., Yo M., Komatsu N., Hiratsuka T., Kogure T., Hoshida T., Goshima N., Matsuda M., Miyoshi H., Miyawaki A.:
“Genetically Encoded Tools for Optical Dissection of the Mammalian Cell Cycle.”
Molecular Cell 68: 626-640. (2017) - Miyawaki A.:
Roger Y. Tsien (1952-2016).”
Cell 167 (2): 298-300. (2016) - Miyawaki A.:
"Exploiting the cyanobacterial light-harvesting machinery for developing fluorescent probes."
Nature Methods, 13 (9): 729-730 (2016).
10.1038/nmeth.3983 - Hama H, Hioki H, Namiki K, Hoshida T, Kurokawa H, Ishidate F, Kaneko T, Akagi T, Saito T, Saido T, Miyawaki A.:
"ScaleS: an optical clearing palette for biological imaging."
Nature Neuroscience, 18 (10): 1518-1529 (2015).
10.1038/nn.4107 - Miyawaki A, Niino Y.:
"Molecular spies for bioimaging - Fluorescent protein-based probes."
Molecular Cell, 48 (4): 632–643 (2015).
10.1016/j.molcel.2015.03.002
News & Media
-
-
Real x-ray vision: see-through brains ready for study
-

-
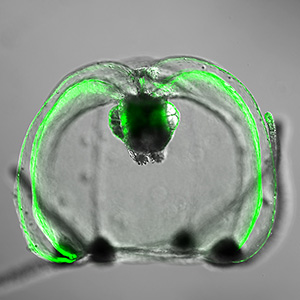
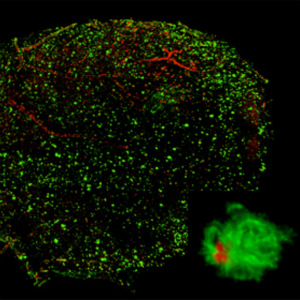
Cells target giant protein crystals for degradation
-
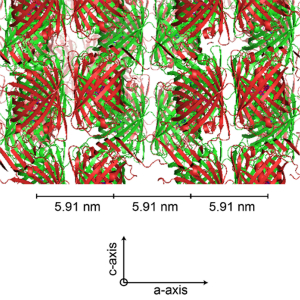
-
Cells target giant protein crystals for degradation
-
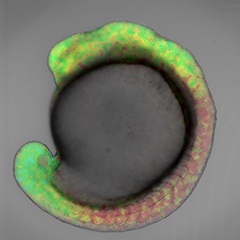
-
Retinoic Acid Gradient Visualized for the First Time in an Embryo
-

-
New chemical reagent turns mouse brain transparent
-

-
Effects of anti-cancer drugs reassessed using fluorescence imaging technology
-

-
Embryogenesis propagation and differentiation patterns of live fish exposed to full view
-

-
Differential silhouetting of the cell cycle specific to either S/G2/M a new visualizing technology of the cellular proliferation process
-

-
Elucidating the molecular mechanism of photochromism of a fluorescent protein 'Drompa' with NMR analysis of its dark-structure, instead of X-ray structural analysis
-

-
Color-coded mouse genes reveal hidden story on cell cycle powerful new tool enables a pictorial roadmap of cell transitions from their earliest stage to functional maturity.
-

-
Color and light to illuminate biochemical pathways
-

-
Discovery of a new fluorescent protein that changes from green to red with ultraviolet light: Development of a reliable basic research tool that can mark cells with light.
-

-
Discovery of the information transmitting mechanism in cells triggered by local stimulation.
-

-
Development of new fluorescent protein that enables the detailed study of live cells: Unknown phenomena inside cells are now visible.
Lab Members
Principal investigator
- Atsushi Miyawaki
- Team Director
Core members
- Peter Greimel
- Senior Research Scientist
- Ryoko Ando
- Research Scientist
- Asako Sakaue
- Research Scientist
- Takashi Fukano
- Research Scientist
- Hiroyuki Kawano
- Research Scientist
- Masahiko Hirano
- Research Scientist
- Satoshi Shimozono
- Research Scientist
- Hiroyuki Katayama
- Research Scientist
- Takayuki Michikawa
- Research Scientist
- Mayu Sugiyama
- Research Scientist
- Hiroshi Kurokawa
- Research Scientist
- Yusuke Niino
- Research Scientist
- Kana Namiki
- Research Scientist
- Naoki Komatsu
- Research Scientist
- Satoshi Iwano
- Special Postdoctoral Researcher
- Hiroshi Hama
- Research Specialist
- Keiko Shoda
- Technical Staff I
- Takako Kogure
- Technical Staff I
- Rie Ito
- Technical Staff I
- Tetsushi Hoshida
- Technical Staff I
- Asako Tosaki
- Technical Staff Ⅱ
- Misaki Kubota
- Technical Staff II